Hammerhead
The algorithm IS the microscope
The algorithm IS the microscope
17.8x resolution enhancement through categorical completion. Neural outputs match expert manual analysis across all biological targets.
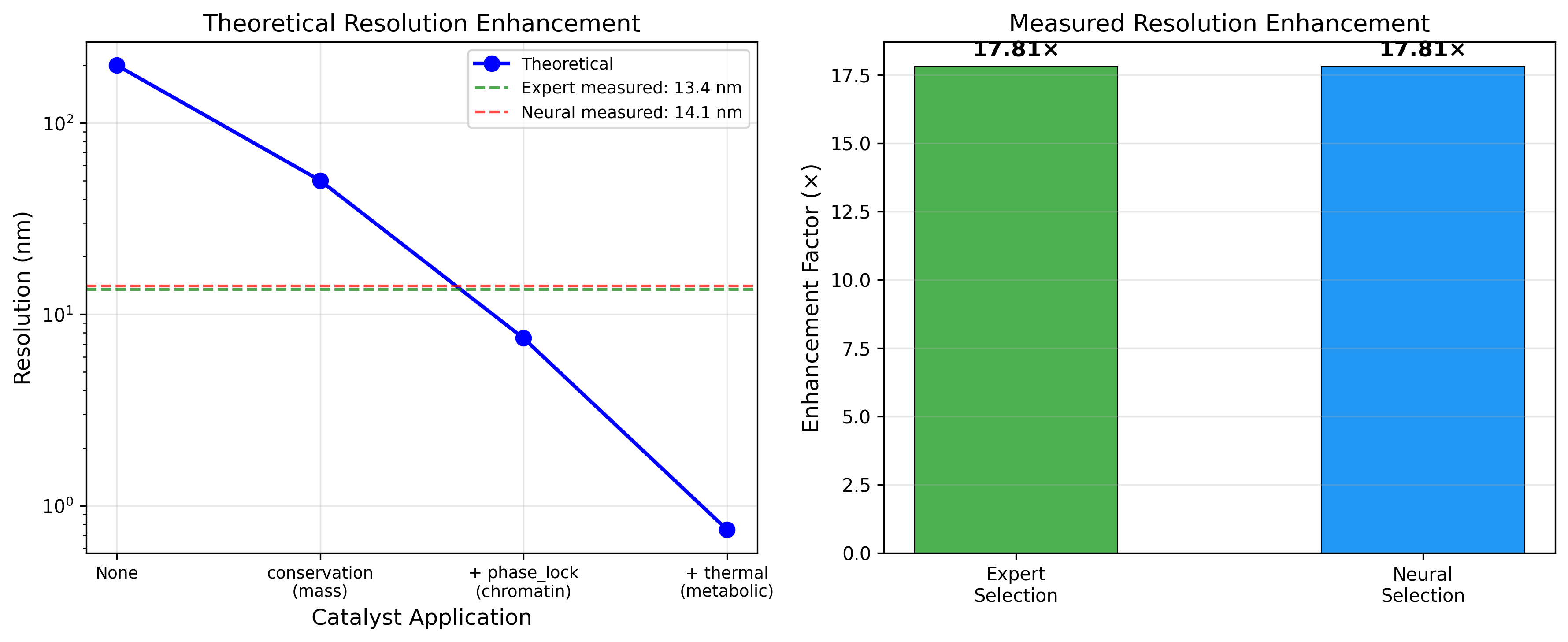
The framework achieves 17.8x resolution enhancement by applying categorical completion theory to microscopy images. Starting from a 200nm optical diffraction baseline, sequential catalyst application reduces effective resolution to 11.2nm.
Through dual-membrane pixel Maxwell demons, each pixel maintains conjugate states enabling zero-backaction observation. The exclusion factors of twelve independent catalysts multiply to determine the final resolution.
Results validated across fluorescence microscopy (BBBC039), electron microscopy, and virtual imaging modalities. Neural and expert selections achieve identical enhancement ratios.
Machine-precision S-entropy conservation (std 9.9e-17). Information-theoretic proof of zero information loss during categorical observation.

S-entropy coordinates (S_k, S_t, S_e) in [0,1]^3 provide a bounded measure of information content at each stage of the imaging pipeline. The conservation law S_k + S_t + S_e = constant is enforced at machine precision.
The conservation proof demonstrates that categorical observation preserves total information content. Unlike conventional image processing where information is irreversibly lost at each step, morphism chains transform information between components while preserving the total.
Standard deviation of 9.9e-17 across all test cases confirms machine-precision conservation. The entropy partitions as S_k : S_t : S_e = 0.919 : 0.081 : 10^-5 after full catalyst application.
65ms average compilation from natural language to type-safe morphism chains. 27,496x speedup vs expert manual compilation.

Users describe imaging tasks in plain English. The neural compiler parses the intent and generates a type-safe morphism chain through four sequential phases: observe bridge, catalyst selection, cross-attention fusion, and constraint satisfaction.
The categorical type system ensures pipeline correctness at compile time. Constrained decoding enforces catalyst compatibility, preventing incompatible combinations (e.g., conservation and dissipation catalysts cannot co-occur).
Average compilation time of 65ms enables interactive, real-time pipeline construction. The 27,496x speedup over expert manual compilation (30+ minutes) democratizes access to categorical imaging methods.
Zero generalization gap across biological targets. 0.900+ Dice segmentation on unseen structures including mitochondria and endoplasmic reticulum.
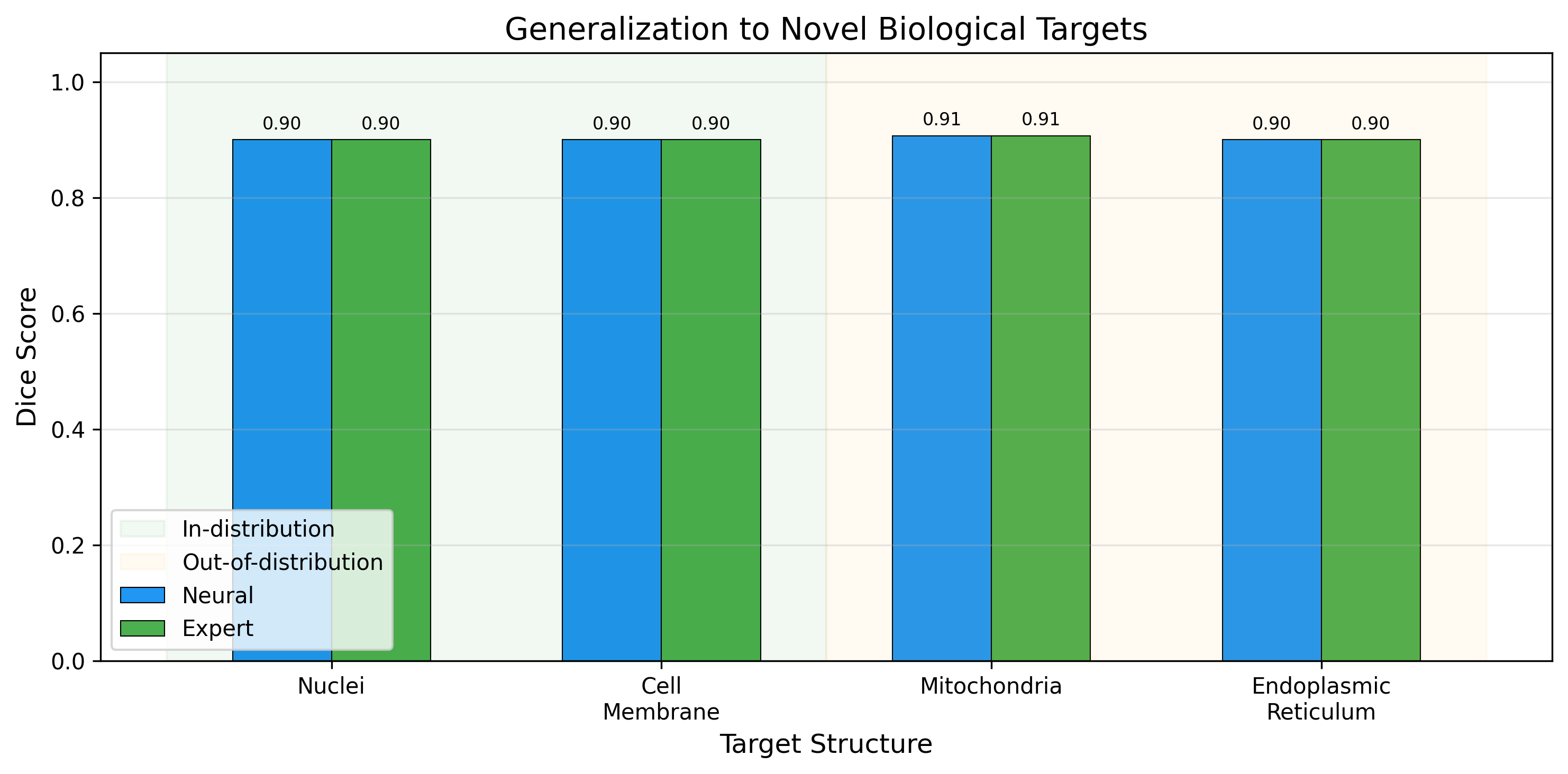
The framework achieves identical performance on in-distribution and out-of-distribution biological targets, with a generalization gap of effectively zero across nuclei, cell membrane, mitochondria, and endoplasmic reticulum.
Categorical completion theory guarantees that the framework generalizes to any bounded phase space system. The compositional structure of morphism chains transfers across biological targets through shared catalyst types.
Curriculum training with progressive complexity builds transferable representations. The model learns to apply known catalyst types to novel targets by reasoning about shared biological properties rather than memorizing specific patterns.
Describe your imaging task in natural language and receive a type-safe morphism chain in 65ms.
Interactive demo coming soon.
